Parkland Center for Clinical Innovation Expands Opportunities for Women with Data Science and Technology Summer Internship Program
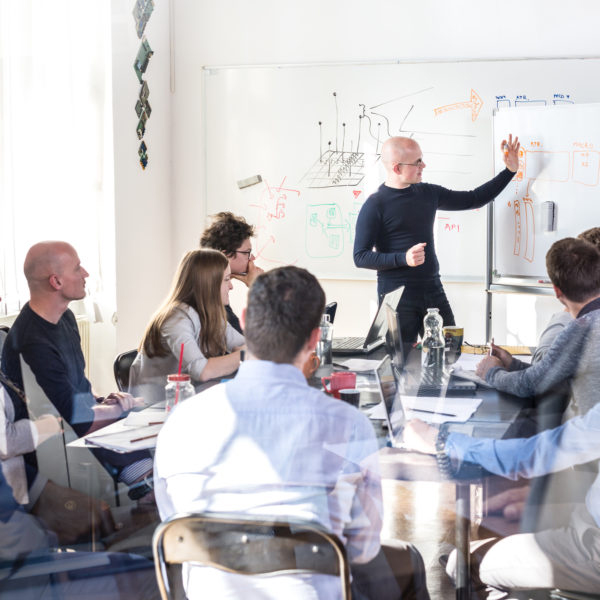
DALLAS – Parkland Center for Clinical Innovation (PCCI), improving healthcare in our communities with advanced analytics and artificial intelligence, recognizes the importance of a STEM education. Offering opportunities to women interested in data science is particularly crucial, which is the mission of PCCI’s summer internship program.
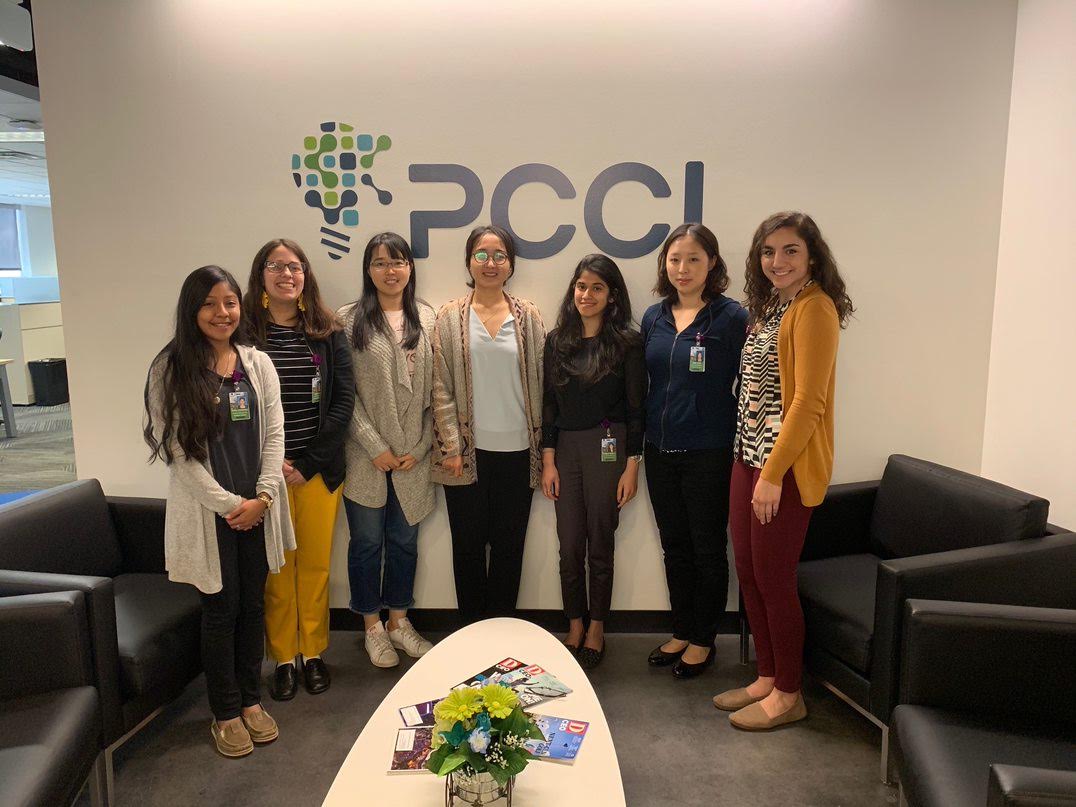
PCCI’s Women in Data Science and Technology Summer Internship, in collaboration with Southern Methodist University’s (SMU) Statistics Department, is one of the most prestigious internship programs in North Texas with a mission to expand opportunities for women in an industry that significantly lacks gender diversity.
The seven women participating in PCCI’s Women in Data Science and Technology Summer Internship program include high school, college and graduate students from Dallas Independent School District high schools, SMU’s Statistics Department as well as students from the University of Texas at Dallas and Creighton University.
The program’s interns will be immersed in PCCI’s daily work where they will directly experience the organization’s innovative healthcare and Non Medical Drivers of Health programs. The students will also have hands-on exposure to the practical applications of analytics, computing and data science.
“The Women in Data Science and Technology Summer Internship program is a rigorous and meaningful path that demonstrates to women what to expect and how to enter the technology market,” Steve Miff, PhD, President and CEO of PCCI. “Because of the important and valuable contributions from organizations such as SMU’s Statistics Department, we are able to place women side-by-side with clinical and data science experts where they can hone their programming and analytics skills within an atmosphere of mentorship and advancement.”
PCCI celebrates diversity and inclusion with a workforce that includes 54 percent women with 30 percent of its employees representing various ethnicities and communities from around the world. As an example of PCCI’s successful commitment to diversity, the Dallas Business Journal recently named Priyanka Kharat, PCCI’s Vice President, Data Engineering and Machine Learning, as a 2019 Women in Technology honoree.
PCCI’s Women in Data Science and Technology Summer Internship program is currently underway and will conclude in mid-August with a presentation program for their PCCI mentors showcasing the impact their projects are having on the Dallas community and Parkland Health & Hospital System.
About Parkland Center for Clinical Innovation
Parkland Center for Clinical Innovation (PCCI) is an independent, not-for-profit, healthcare intelligence organization affiliated with Parkland Health & Hospital System. PCCI focuses on creating connected communities through data science and cutting-edge technologies like machine learning. PCCI combines extensive clinical expertise with advanced analytics and artificial intelligence to enable the delivery of patient-centric precision medicine at the point of care.
###
Parkland Health & Hospital System, Department of Corporate Communications
5200 Harry Hines Blvd., Dallas TX 75235, 469-419-4400
D CEO Healthcare: North Texas Healthcare Innovation: Fighting Fungus, Sharing Data, Virtual Reality, and Radio Therapy
The North Texas healthcare market is constantly changing and innovating, bringing original ideas, techniques, and technology to patients in the region. We decided to check in with a startup, a nonprofit, a provider and an academic institution who are on the leading edge of healthcare innovation, and they told us about the latest procedures, techniques, software, and technology making a difference for patients.
Continue readingMy PCCI Internship – Synthetic Data Project
As my internship at Parkland Center for Clinical Innovation (PCCI) comes to an end, it feels great to look back and ponder over what I had the opportunity to work on, achieve and experience over the past three months. I arrived at PCCI with high expectations and am happy to say that I wasn’t disappointed. The project I worked on is called “Synthetic Data.” As the name suggests, the goal of the project is to create synthetic data from real medical datasets.
Why do we need synthetic medical data sets?
Real medical data is expensive and seldom released for research due to various privacy issues connected to it. Regulations exist because by looking at the medical data, a hacker could identify the name of a patient, thereby gaining access to sensitive information. The synthetic data project at PCCI aims to alleviate these problems by creating synthetic datasets, which are as close to the real medical datasets as possible without compromising a patient’s privacy.
Generative Machine Learning Algorithms and Challenges
Generative machine learning algorithms, specifically, Generative Adversarial Networks (GANs), proposed in 2014 [1] were used in this project. GANs have gained huge popularity within the machine learning community with a wide variety of GAN models being proposed.
There were quite a few challenges along the way in realizing the goal of synthetic data generation. First, the proposed GAN model has not been applied to real medical datasets before us, as it was mainly designed for image generation tasks. It also tends to not perform well with different modalities of data, which are naturally present in a real medical dataset. Modifying the network to work with real medical data or modifying the data (mostly getting it into a single modality) for it to work with GANs was a major challenge.
Second, there is an explosion in the number of GAN-based architectures being proposed and thus coming up with a novel architecture is a huge challenge in itself. After a lot of deliberation, we came up with an approach that allows us to incorporate domain knowledge into the GAN architecture. Below were our three possible approaches:
- Advice on just the discriminator.
- Advice on just the generator.
- Advice on improving the zero-sum game between the discriminator and the generator.
Improving the Zero-sum Game
Taking the third approach, we decided to incorporate reconstruction error [2] into the discriminator and the generator loss functions. The simple intuition is this: train the network with a mini-batch of data and generate the synthetic counterpart for each mini-batch. Since reconstruction errors, as the name suggests, measure the errors between the real data and the “reconstructed” data, adding this to the loss functions can penalize either the discriminator or the generator depending on which performs the worse (i.e. has a higher loss difference) with respect to the mini-batch in question.
Initial Experiments Using the MIMIC III Dataset
For our initial experiments, we made use of the MIMIC III dataset [3], which is a dataset incorporating clinically relevant data for all admissions to an ICU at the Beth Israel Deaconess Medical Center between 2001 and 2012. Figure 2 shows the selected features.

Experiments Using GAN-based Methods
We then ran a couple of experiments. First, we used three different GAN-based methods, without the reconstruction error, to see how the network performed with the defined loss functions. Figure 3 shows the original and the generated data as obtained from these networks.

An Imbalanced Dataset
The original and synthetic datasets were then used to train a machine learning model. For the synthetic data to be useful, it needs to be as close to the real data distribution as possible, which should be captured by a machine learning classification model. Our dataset was highly imbalanced. We were predicting mortality rate and since a majority of patients come out of the ICU alive, we had nearly a 90%-10% split between negative examples i.e. patients who are alive after the ICU treatment and positive examples i.e. patients who die in the ICU. We used a cost-sensitive support vector machine classifier, constructed for such imbalanced data, to report the F1 score and the area under the curve (AUC-ROC) in Figure 4.

As it can be seen, the results proved our hypothesis that real healthcare data was going to be a challenge for techniques, like GANs, which rely on many samples of very predictable data types since healthcare data tends to be more diverse and are difficult to compose into higher-order features. (Credit: David Watkins, my supervisor). We then used the reconstruction error to “indirectly” capture the relationships between the data points and used “real” hospital data to test on and create a synthetic dataset from it. The work was currently still in progress at the time this blog was written.
Working at PCCI
The work culture at PCCI, in my opinion, towers above other places. The work hours are flexible, the team’s ethics and bonding are strong and people are always willing to help you regardless of their schedule. The company truly values its employees and creates a work environment where every employee gives his/her best. You will never feel out of place (not even on the first day) as all your tasks are defined and everyone is so welcoming. A great thing about PCCI is the absence of an implicit hierarchy. Everyone from the CEO to your respective manager(s) (thank you Albert Karam) is always accessible. I never felt any different than a PCCI employee and this says a lot about the values of the company and how these values are being nurtured by PCCI’s CEO Steve Miff and all employees of this amazing organization.
Another important quality about PCCI is that it values and encourages all feedback that any employee may have and any grievances are then actually addressed. I am proud to say that Steve himself makes sure that any such issues are addressed.
Nothing is perfect and PCCI has a few areas where it can improve. One area of improvement is getting access to the real data. This is currently a very is a slow process, which makes sense as it is sensitive medical information of real patients, but it can be sped up. Another area of improvement, that I have actually raised to Steve during a meeting is that PCCI should focus on publishing research papers. It is a company that is capable of doing amazing research and has access to real medical datasets that are difficult to find. I hope PCCI becomes more active in this regard.
What PCCI does is super important to the community and it makes sure that all the employees realize this fact. Creating an impact in the real world, on real lives is a great morale booster for anyone and since PCCI values are centered around this motto, working here has been a great experience. It’s been a pleasure interning at PCCI and I am happy to be taking fond memories with me back to school.
Learn more about PCCI’s careers, or stay-up-to-date with our recent news by following us on Facebook, Twitter and LinkedIn!
References:
[1] Goodfellow, Ian, et al. “Generative adversarial nets.” Advances in neural information processing systems. 2014.
[2] Borji, Ali. “Pros and Cons of GAN Evaluation Measures.” arXiv preprint 2018.
[3] Johnson, Alistair EW, et al. “MIMIC-III, a freely accessible critical care database.” Nature 2016.
My Summer as a Data Science Intern at PCCI
For the short duration of returning to my hometown Dallas for the summer, I’ve been interning at Parkland Center for Clinical Innovation (PCCI) as a Data Science Intern. During my interview with Albert and Vikas, we discussed some issues with the representation of data in the current healthcare system. Hospitals use different coding systems in their electronic medical records (EMRs), making communication between hospitals and care providers difficult. A while ago, a new health data standard called FHIR (Fast Healthcare Interoperability Resource, pronounced “fire”) was proposed. My project this summer aimed at identifying whether data could be easily transformed into the new FHIR format, carrying out the transformation, and creating predictive models using the new FHIR data.
Situated on the 11th floor of the building, PCCI is a very chill place to work. Quiet spaces are easily found at desks and conference rooms scattered around the office. As an intern, I sit on the “Intern Island” with (usually) 6 other interns. I like this space because we get two monitors and a Lenovo Thinkpad.

As for work, each PCCI project usually consists of one project manager, a clinical expert, and a data scientist. The intern projects are no different; Aaron was the FHIR Project Manager Intern, and Mila was the FHIR Clinical Intern. Both had important but separate duties that helped our project succeed.
As the Data Science Intern on the FHIR project, I was responsible for first converting the data into FHIR resources. This involved bringing back Java knowledge from several years ago! There were definitely some issues figuring out how to add the right dependencies because Java can get complicated very quickly. A few days were spent just trying to get oriented with Java and Eclipse, and making sure all the necessary packages for FHIR were installed.
We were working with two years of data. This roughly translates into 27 million (!) vitals and 17 million labs, and each vital and lab was converted into its own separate file. I quickly realized that there would be no space on my laptop to hold all of these files, so we decided to enlist the help of Microsoft Azure. With Azure, the task became less difficult, but still, the hardest part of my summer was working with such huge numbers of files.
Caught up in the huge task of transforming vast amounts of data to FHIR resources, I left very little time in my internship to work on actual data science. Out of the approximately 13 weeks total, about six weeks were spent converting the table format EMR data into FHIR resources, five weeks were spent on parsing the FHIR resources into a format for machine learning, and the remaining two weeks were dedicated to model building. Reflecting back, I would definitely work harder to cut short the resource conversion in favor of more time for data science.

As a Data Science Intern at PCCI, you have the freedom to work in any language you want; the full-time Data Science team is very evenly divided between R and Python. There’s also a lot of freedom in dictating which path your project will go. Your supervisor will point you in a very general direction of where to go and state goals and expectations, but is otherwise very lenient!
Don’t be shy about asking around people for advice and help, even if they’re not on your project team! Even though most people are busy with various meetings, they will gladly schedule a 30-minute or even hour-long block to discuss your project privately with you.
When presenting your project, whether it’s a progress update or final presentation, expect multiple questions from the audience. It’s not that they want to quiz you on your knowledge and preparation on your project, but because they’re genuinely curious and care about understanding what you’re doing over the summer.
A mandatory 30-minute lunch is required every day. I recommended bringing lunches that can stay in the fridge for several days (like salad) or not bringing anything because there are often team lunches and random outings during the day. Occasionally there’s leftover pizza or sandwiches from lunch meetings in the big conference room or leftover burritos from breakfast.
I enjoy the diverse atmosphere at PCCI the most. The three teams: Data Science, Project Management, and Clinical teams collaborate and work together so well. It’s a very fluid system. A data scientist with a question about the best intervention methods for patients with diabetes can easily walk over to a clinical team member and get an answer within minutes. Despite being employed as a data scientist, you have access to an entire host of medical knowledge from the clinical team and connections from the project management team.
My biggest takeaway from this internship is learning about long-term time management and collaboration. Manage your time well and you’ll be able to at least touch on everything you wanted to learn during your internship. Collaborate with as many people as you can, so not only can you learn so much more but also gain friends and connections while doing so.
The Increasing Importance of Non Medical Drivers of Health
IMPACT ON HEALTH OUTCOMES
Over the last few years, it has been very clear from research that Non Medical Drivers of Health (NMDOH) variables have a major impact on health outcomes. It is estimated that close to 80% of health outcomes are impacted by NMDOH. With the rise of population-based risk contracts in both the commercial and government sector, it is essential for both providers and payers to collaborate in the identification of best practices to address these NMDOH variables. This is especially relevant as providers such as hospitals assume greater risk in arrangements with plans throughout the country such as Accountable Care Organizations (ACO) and bundled payments.
NATIONAL INTEREST AND PROGRESS
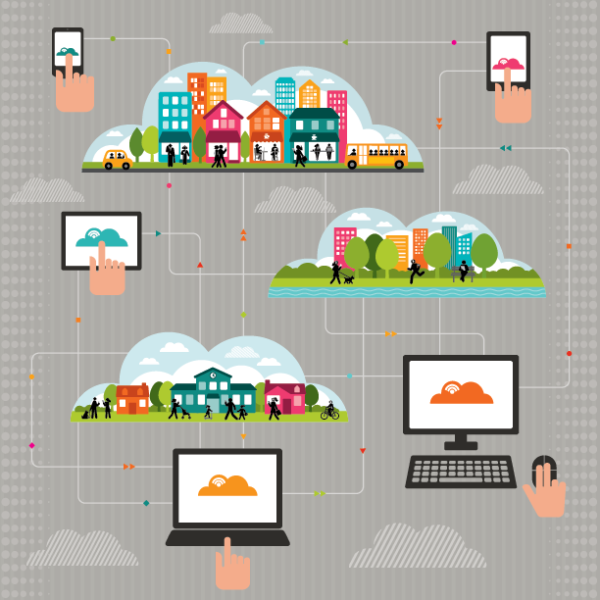
Many national associations such as the American Hospital Association (AHA) and America’s Essential Hospitals have developed resources and launched learning collaboratives for hospitals and health systems to address these variables such as food insecurity, housing, and transportation. Health system innovation and care-redesign models driven by organizations such as Healthbox and AVIA have launched collaboratives and forums to educate and address NMDOH initiatives. The May 3, 2018, Healthbox forum discussion on “Challenging the Status Quo of Social Determinants” visually captured the opportunities and challenges ahead into one image (Figure 1):

These variables have always been a focus of many health systems in terms of articulating their benefit to the community, but now they have particular importance given the rise of more population risk contracts.
Several major barriers have impeded the industry’s progress in addressing NMDOH variables: funding and regulations. Fortunately, we have begun to see opportunities in both areas emerge in 2018!
MEDICARE UPDATES AND THE BENEFITS OF Non Medical Drivers of Health DATA
Medicare Advantage (MA) has a regulation titled “Uniformity Standard” that requires all of the plan’s benefits, including cost-sharing, be the same for all plan enrollees. On April 2, 2018, the Centers for Medicare & Medicaid Services (CMS) outlined several widespread changes in this regulation that both providers and plans have advocated for over the last several years in their 2019 Medicare Advantage Call Letter. CMS expanded the flexibility of lifting the uniformity of supplemental benefit to allow different segments of an MA plan to offer specific benefits to a targeted population like diabetics. This can begin in CY 2019 (January 1, 2019) after the plan designs are approved by CMS. An example could be reduced cost sharing for foot or eye exams. In their official bids that were submitted by the June 4, 2018 deadline, the MA plans can include any of these supplemental benefit elements. Hopefully, providers will see many of the plans deciding to include these additional benefits in their MA bids to address the NMDOH variables.
Additionally, in the Bipartisan Budget Act (BBA) that was passed in early 2018, Congress has taken it further by extending the lifting of the uniformity of the supplemental benefits to all chronically ill members of the MA plans effective January 1, 2020. This reinforces the need for us to gain valuable lessons during 2019 in order to determine what works and what doesn’t before it is transitioned to a broader population.
The Chronic Care Act of 2018 extended the Center for Medicare & Medicaid Innovation’s (CMMI) Valued-Based Insurance Design Model to all 50 states in 2020. This model was launched in 2017 to allow Medicare Advantage plans to offer supplemental benefits and reduced cost-sharing to seven conditions including Coronary Artery Disease or Congestive Heart Failure. The model focuses on four approaches:
- Reduced Cost Sharing for High-Value Services
- Reduced Cost Sharing for High-Value Providers
- Reduced Cost Sharing for enrollees participating in disease management
- Coverage of additional supplemental benefits such as transport or meal delivery
The creation of more supplemental benefits will enhance the quality of services we provide for our patients especially in terms of addressing the NMDOH. Encouraging the inclusion of these targeted supplemental benefits will allow us to partner with payers to improve the health of the country in a more innovative way.
ADDRESSING NMDOH WITH HEALTHCARE PROVIDERS AND COMMUNITY RESOURCES
At PCCI, we have been directly involved in national and state-driven education forums, presentations, and roundtables directed to design and deploy local models for the Connected Communities of Care program (previously known as the Information Exchange Portal) that bring together providers, payers, philanthropic organizations, community-based organizations (CBO), and local/state government entities. While most markets continue to be in a learning mode, significant and tangible activities are being initiated in a number of municipalities, including Dallas, Raleigh-Durham, Louisville, Detroit, Chicago, Phoenix, Salt Lake City, as well as across whole regions. For example, North Carolina recently requested proposals for the development of a North Carolina Resource Platform via the Foundation for Health Leadership & Innovation. The goal of this multi-year program is to connect over 3,000 statewide community-based organizations via technology, and facilitate NMDOH. This will be completed through a programmatic coordination of referrals between healthcare providers and community resources to comprehensively identify and address the needs of individuals across the state. On a broader level, the Accountable Health Communities Model deployed in 2017 is engaging 31 organizations across the country to address a critical gap between clinical care and community services in the current healthcare delivery system. This is being done by testing the process of systematically identifying and addressing the health-related social needs of Medicare and Medicaid beneficiaries through screening, referral, and community navigation services to see if it will impact healthcare costs and reduce healthcare utilization.
SUCCESS IN SIX TRACKS
Our experience over the last five years across Dallas tells us that models will need to address six tracks to be successful: Governance, Legal, Technology, Clinical Workflows, CBO Workflows, and Sustainability (Figure 2). The maturity and evolution of the models need to develop and be staged within a multi-year deployment framework (concentric circles in Figure 2 represent the progression and evolution of the model with outer circles representing mature and more sophisticated models).

There is also a critical upfront readiness and deployment/implementation assessment that is important in order to stage the deployment of a Connected Community of Care program. This broad representation of the community’s fabric is critical to ensure that:
- A community is ready to undertake the operational and financial requirements associated with deploying a Connected Communities of Care program
- The healthcare and social needs of the community are at the forefront of the customized design of the platform (something most for-profit technology vendors offering an out-of-the-box solution either cannot do or fail to do properly)
- The design is sufficiently flexible to adjust as the healthcare or social needs of the community change
Addressing NMDOH is finally moving from a “buzz” word to implementation pilots. While we talked a lot about population health over the last 10 years, doing population health without a truly engaged and “Connected Community of Care” is like focusing on rescuing people from drowning in a river vs. building a bridge so they can cross it safely. As we continue this journey, let us make sure we build a bridge that adapts to the needs of each community and has emerging local and national models of care to ensure sustainability. We don’t want to end up with a bridge like the Choluteca Bridge in Honduras, connecting nothing to nowhere.
Acknowledgments: Valinda Rutledge, PCCI Executive Advisor and Lindsey Nace, PCCI Marketing and Communications have contributed to this article.
Stay up-to-date with PCCI’s data science work by checking our recent news and follow us on Facebook, Twitter and LinkedIn!
Hired at “I wrote you a code”
First Question, Lasting Impression
How often did you get the question (or have asked it yourself) during an interview: “Why are you interested in our organization?” Simple, standard, mundane – on the surface, some might even call the question un-inspiring. I disagree. It tells me right off the bat how much effort the candidate put into learning about the Parkland Center for Clinical Innovation, our organization, our work, our team and how they synthesized and interpreted the information. I can tell within the first two minutes how interested I will be for the next 28. Regardless of the level of experience, I’m way too often disappointed by the response.
Using Python to Create a Sentiment Analysis
Recently, I was blown away! I asked the same “boring” question to a candidate interviewing for an entry data science position at PCCI. As soon as I finished the question, his eyes lit up and he quickly pulled out a document from his bag. With great enthusiasm, he replied:

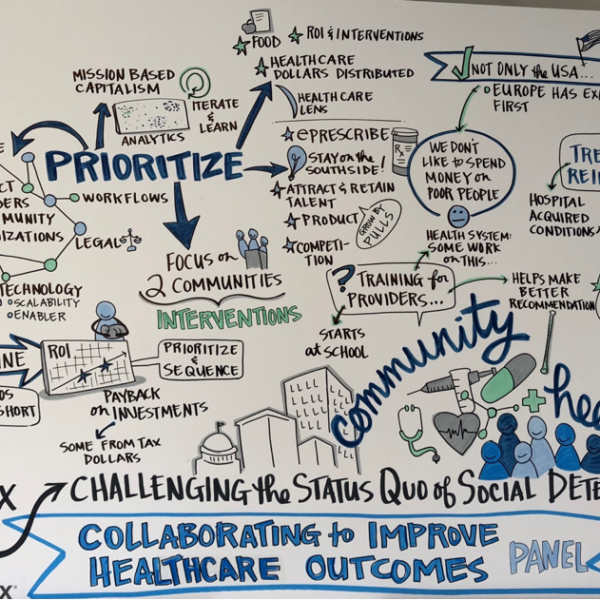
“In addition to my own research, I wanted to know what others are saying and feeling about PCCI. So, I wrote code in Python to create a Twitter sentiment analysis. I used it to create a word cloud and analyzed it to see if the keywords match my passion and interpretation of my own research. These four words really resonated with me because … I also wanted to understand PCCI’s reach and brand recognition, so I analyzed the top 10 famous people and companies talking about PCCI. I was impressed to see @HarvardBiz, @washingtonpost, @NIH, @HHSGov, etc, but most importantly to see @KirkDBorne. He’s so influential. Finally, the outputs of the Sentiment Count Plot Analysis and the Sentiment Subjectivity Distribution reconfirmed that this is a great place and the place I want to be.”
A Match Made in Data Science
I know I’m a geek at heart and this answer resonated with me more than it would with most (did I mention earlier how important it is to know your audience and their interests when answering a question?), but regardless of what approach you take, this is how you do it!
Stay up-to-date with PCCI’s data science work by checking our recent news and follow us on Facebook, Twitter and LinkedIn!